Difference between revisions of "Support:Documents:Examples:Estimate Change of Neurotransmitter"
| Line 8: | Line 8: | ||
[[Image:Model.jpg]] | [[Image:Model.jpg]] | ||
| − | This model can be described by the differential equations: | + | This model can be described by the differential equations: <math>B_en</math> |
dF/dt = K<sub>1</sub>C<sub>p</sub> - K<sub>2</sub>F - K<sub>on</sub>[B<sub>max</sub> - B - B2 - B<sup>en</sup>]F + k<sub>off</sub>B | dF/dt = K<sub>1</sub>C<sub>p</sub> - K<sub>2</sub>F - K<sub>on</sub>[B<sub>max</sub> - B - B2 - B<sup>en</sup>]F + k<sub>off</sub>B | ||
Revision as of 20:40, 11 March 2009
Model of Neurotransmitter PET (ntPET)
Overview
The changes of endogenous neurotransmitter by PET scanning after a stimulus have been proved with different approaches. In stead of detecting the increase or decrease of a neurotransmitter, it is important to characterize the temporal change of a neurotransmitter after a stimulus. Morris proposed the new technique (called ntPET) for capturing the dynamic changes of a neurotransmitter after a stimulus and demonstrated the ability of this method to reconstruct the temporal characteristics of an enhance in neurotransmitter concentration.
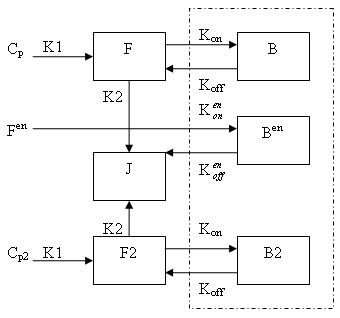
Generally, there are two separate injections of tracer for ntPET: one for the rest condition (without any stimuli) and the other for the activation condition (after a stimulus). Therefore, the model can be described as:
This model can be described by the differential equations: <math>B_en</math>
dF/dt = K1Cp - K2F - Kon[Bmax - B - B2 - Ben]F + koffB
dB/dt = Kon[Bmax - B - B2 - Ben]F - koffB
dF2/dt = K1Cp2 - K2F2 - Kon[Bmax - B - B2 - Ben]F2 + koffB2
dB2/dt = Kon[Bmax - B - B2 - Ben]F2 - koffB2
dBen/dt = Kenon[Bmax - B - B2 - Ben]Fen -kenoffBen
Cp is the plasma concentration of tracer at the first injection (the "rest" condition).
F and B are free (unbound) and bound molar concentrations of tracer after the first injection
Cp2 is the plasma concentration of tracer at the second injection (the "activation" condition).
F2 and B2 are free (unbound) and bound molar concentrations of tracer after the second injection.
Fen and Ben are free and bound molar concentrations of a neurotransmitter released by endogenous ligand after a stimulus.
Bmax is the maximum number of receptors that can be bound by neurotransmitter or tracer.
Bmax - B - B2 - Ben is the concentration of available receptors. This also indicates that binding is saturable.
K1, K2, Kon, Koff, Kenon and Kenoff are rate constants.
Implementing the Compartment Model
Create a new model for ntPET
Step 1. Create the compartemt model
cm = compartmentModel;
%Set scan time of the whole scan
inj2Delay = 240; % min between 1st and 2nd inj
disp('Define one long study: inj 1 = baseline, inj 2 = activation');
scantInj1 = [[0:0.1:0.9 1:0.25:1.75 2:0.5:4.5 5:1:59]' [0.1:0.1:1 1.25:0.25:2 2.5:0.5:5 6:1:60]'];
scantInj2 = scantInj1 + inj2Delay;
scant = [scantInj1; scantInj2];
deltaT = scant(:,2) - scant(:,1);
tmid = 0.5*(scant(:,1) + scant(:,2));
cm = set(cm, 'ScanTime', scant);
%Set parameters of the model
cm = addParameter(cm, 'PV', 1 );
cm = addParameter(cm, 'Fv', 0);
cm = addParameter(cm, 'sa', 1); % uCi/pmol
cm = addParameter(cm, 'dk', 0); % assume data are decay-corrected
cm = addParameter(cm, 'k1', 0.9);
cm = addParameter(cm, 'k2', 0.4);
cm = addParameter(cm, 'kon' , 0.01);
cm = addParameter(cm, 'koff', 0.14);
cm = addParameter(cm, 'k2ref', 0.3);
cm = addParameter(cm, 'Bmax', 80);
cm = addParameter(cm, 'konEN', 0.25);
cm = addParameter(cm, 'koffEN', 25);
%model for baseline condition:
cm = addCompartment(cm, 'F');
cm = addCompartment(cm, 'B', 'Bmax');
cm = addCompartment(cm, 'Junk');
%model for activation condition:
cm = addCompartment(cm, 'F2');
cm = addCompartment(cm, 'B2', 'Bmax');
cm = addCompartment(cm, 'B^{EN}', 'Bmax');
Step 2. Set the input function for two injections
% Create input functions for two injections
t = 0:0.05:305; % coverage out to 6+ h
Cp1Data = fengInput([2 0.5 8.5 0.22 0.2 -4. -0.1 -0.02], t);
Cp2Data = fengInput([2 inj2Delay+0.5 8.5 0.22 0.2 -4. -0.1 -0.02], t);
ppCp1 = spline(t, Cp1Data);
ppCp2 = spline(t, Cp2Data);
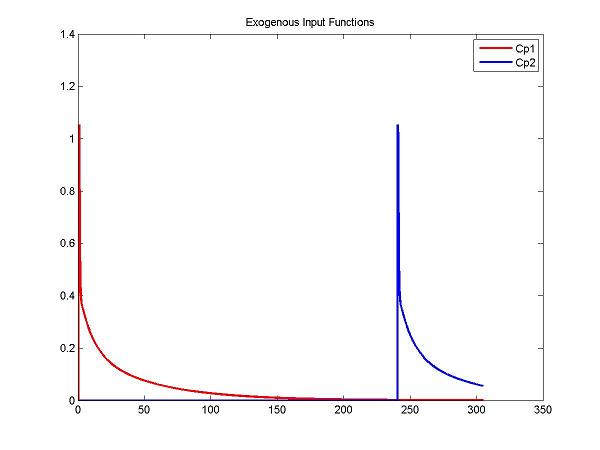
figure; plot(t, ppval(ppCp1, t), 'r', t, ppval(ppCp2,t), 'b', 'LineWidth', 2); title('Exogenous Input Functions'); legend('Cp1', 'Cp2')
cm = addInput(cm, 'C_p', 'sa', 'dk', 'ppval', ppCp1);
cm = addInput(cm, 'C_p2', 'sa', 'dk', 'ppval', ppCp2);
Step 3. Create a function described the change of concentrations of the endogenous neurotransmitter. To detect the change, the parameters of the function will be estimated.
The change of concentrations of the endogenous neurotransmitter after a stimulus is described by a gamma variate function. Fen = Basal + Gamma[t-Delay]Alpha exp(-Beta[t-Delay]). Basal is the concentration of the endogenous neurotransmitter without any stimulus. Gamma, Alpha and Beta control the change of concentrations of the endogenous neurotransmitter. Delay is the delay time.
Note: For ntPET, the initial concentration of the endogenous neurotransmitter (i.e., Ben) is not zero. At time=0, dBen/dt=0 and B=0. Under this condition, Ben=BmaxFen/(kenoff/kenon+Fen)
This indicates thtat Ben depends on the Fen. which is equal to the parameter 'Basal' as time=0. Therefore, during each iteration of solving system models or curve fitting, the initial concentration of the endogenous neurotransmitter at current iteration is dependent on the previous estimated parameter 'Basal'.
To consider this specific case, ComKat creates a new version of commands for adding parameter. The below command about 'addParameter' is the new version. Instead of using a vector for all parameters, we add each parameter as a cell array, individually. The settings of initial conditions will be described below.
Now, the current version of ComKat does not support this new style. In order to use this new style, please email us. We will send all required files for you. In the future, we will release a new version of ComKat that supports this new style.
DApar = [50 27 1 0.1 240+10]; % = [Basal Gamma Alpha Beta Delay]
params = {'DAbasal', 'G', 'alpha', 'beta', 'delay'};
for i = 1:length(params),
cm = addParameter(cm, params{i}, DApar(i));
end
cm = addInput(cm, 'F^{EN}', 0, 0, 'ntInput', params);
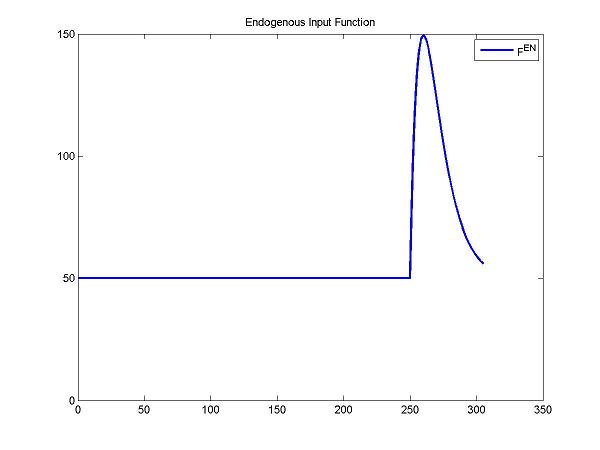
figure; plot(t, ntInput(t,0,params,num2cell(DApar)), 'b', 'LineWidth', 2); title('Endogenous Input Function'); legend('F^{en}');
cm = addSensitivity(cm, params{:}, 'k1', 'k2', 'kon', 'koff');
The endogenous input function is implemented as (create the below function and put in in a file called ntInput.m):
function [Fen, dFendp] = ntInput(t, nsens, pName, pValue, pSensIdx, cm, xtra)
% This function is used to described endogenous dopamine input.
% The model uses this as
% params = {‘DAbasal’, ‘G’, ‘alpha’, ‘beta’, ‘delay’};
% cm = addInput(cm, ‘Fen’, 0, 0, ‘ntInput’, params); % don’t need xparameters
% these are not used in this function – only used here to illustrate
% that comkat passes the parameter name array.
% The names could be used to generalize this so that it could adapt to
% whatever parameters names and order were specified by the user.
% For simplicity, these are hard-coded for this example.
DAbasalName = pName{1};
GName = pName{2};
alphaName = pName{3};
betaName = pName{4};
delayName = pName{5};
% get the values of the named parameters
DAbasal = pValue{1};
G = pValue {2};
alpha = pValue{3};
beta = pValue {4};
delay = pValue {5};
if (nargout > 0),
Fen = DAbasal*ones(size(t));
idx = find(t > delay);
dt = t(idx) - delay;
Fen(idx) = DAbasal + G*(dt.^alpha).*exp(-beta.*dt);
if (nargout > 1),
DAbasalSensIdx = pSensIdx{1}; % location (index) of DA among the sensitivity parameters
GSensIdx = pSensIdx{2};
alphaSensIdx = pSensIdx{3};
betaSensIdx = pSensIdx{4};
delaySensIdx = pSensIdx{5};
% allocate for all sens parameters in model (not just ones relevant to this function)
dFendp = zeros([length(t) nsens]);
dFendp(:, DAbasalSensIdx) = ones(size(t)); % dFen/dDAbasal
if ~isempty(idx),
dFendp(idx, GSensIdx) = (dt).^alpha.*exp(-beta.*(dt)); % dFen/dG
dFendp(idx, alphaSensIdx) = G.*(dt).^alpha.*log(dt).*exp(-beta.*(dt)); % dFen/dalpha
dFendp(idx, betaSensIdx) = -G.*(dt).^(alpha+1).*exp(-beta.*(dt)); % dFen/dbeta
dFendp(idx, delaySensIdx) = -G*alpha*(dt).^(alpha-1).*exp(-beta.*(dt)) + G*beta*(dt).^alpha.*exp(-beta.*(dt)); % dFen/ddelay
end
end
end
Step 4. Set link between each compartment and add output.
%links/output for baseline condition:
cm = addLink(cm, 'L', 'C_p', 'F', 'k1');
cm = addLink(cm, 'K', 'F', 'Junk', 'k2');
cm = addLink(cm, 'K2', 'F', 'B', 'kon');
cm = addLink(cm, 'K', 'B', 'F', 'koff');
disp('Output is sum of radioactivity in free and bound compartments')
WList = {...
'F', 'PV';
'F2', 'PV';
'B', 'PV';
'B2', 'PV'};
XList = {};
cm = addOutput(cm, 'PET Scan', WList, XList);
%links/output for activation condition:
cm = addLink(cm, 'L', 'C_p2', 'F2', 'k1');
cm = addLink(cm, 'K', 'F2', 'Junk', 'k2');
cm = addLink(cm, 'K2', 'F2', 'B2', 'kon');
cm = addLink(cm, 'L2', 'F^{EN}', 'B^{EN}', 'konEN');
cm = addLink(cm, 'K', 'B2', 'F2', 'koff');
cm = addLink(cm, 'K', 'B^{EN}', 'Junk', 'koffEN');
Step 5. Solve the model and generate the output
disp('Properly handle the initial conditions assuming no exogenous ligand initially')
compNames = get(cm, 'CompartmentName');
sensNames = get(cm, 'SensitivityName');
ncomp = get(cm, 'NumberCompartments');
idxBENComp = strmatch('B^{EN}', compNames, 'exact');
idxBasal = strmatch('DA', sensNames);
IC.time = 0;
IC.compartment = cell([ncomp 1]);
IC.compartmentSensitivity = cell([length(compNames) length(sensNames)]);
IC.compartmentSensitivity{idxBENComp, idxBasal} = 'Bmax*(koffEN/konEN) / ((koffEN/konEN + DAbasal)^2)';
IC.compartment{idxBENComp} = 'Bmax*DAbasal/(koffEN/konEN+DAbasal)';
cm = set(cm, 'InitialConditions', IC);
[PET, PETIndex, Output, OutputIndex] = solve(cm);
Bmax = pxEval(cm, 'Bmax');
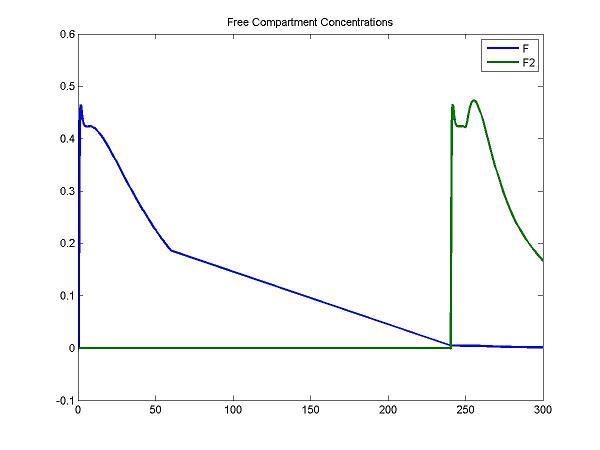
disp('Plot some compartment concentrations to examine model behavior');
figure; idx = [3 6]; plot(Output(:,1), Output(:, idx), 'LineWidth', 2); legend(OutputIndex(idx)); title('Free Compartment Concentrations');
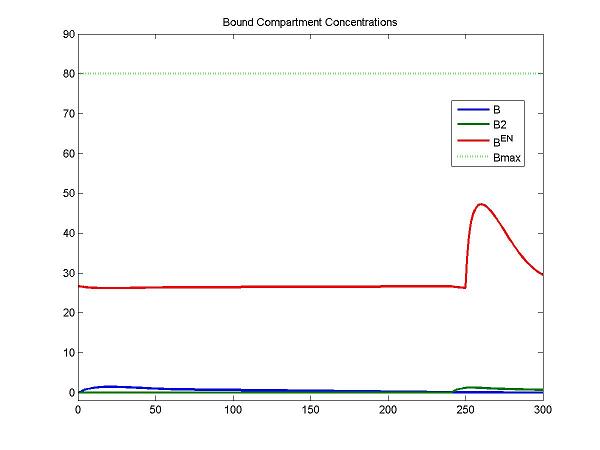
figure; idx = [4 7 8]; plot(Output(:,1), Output(:, idx), [0 Output(end,1)], [Bmax Bmax], 'g:', 'LineWidth', 2);
legend(OutputIndex(idx), 'Bmax'); title('Bound Compartment Concentrations');
Step 6. Fit the simulated data
basal_LB = 30; basal_UB = 70;
gamma_LB = 15; gamma_UB = 50;
alpha_LB = 0.1; alpha_UB = 5;
beta_LB = 0.01; beta_UB = 1;
delay_LB = 240; delay_UB = 300;
k1_LB = 0.01; k1_UB = 3;
k2_LB = 0.01; k2_UB = 1;
kon_LB = 0.001; kon_UB = 0.1;
koff_LB = 0.01; koff_UB = 1;
pinit = [45 18 1 0.01 250 0.1 0.1 0.005 0.1];
cm = set(cm, 'ExperimentalData', PET(:,3));
lb = [basal_LB ; gamma_LB ; alpha_LB ; beta_LB ; delay_LB ; k1_LB ; k2_LB ; kon_LB ; koff_LB];
ub = [basal_UB ; gamma_UB ; alpha_UB ; beta_UB ; delay_UB ; k1_UB ; k2_UB ; kon_UB ; koff_UB];
[pfit, modelfit, resnorm, residual, exitFlag, output] = fit(cm, pinit, lb, ub);
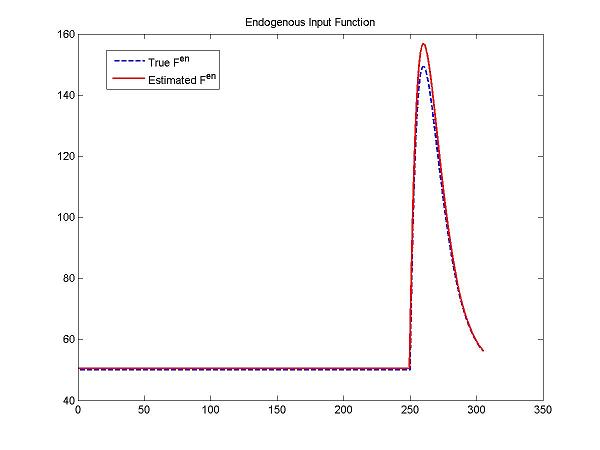
figure; plot(t, ntInput(t,0,params,num2cell(DApar)), 'b--',t, ntInput(t,0,params,num2cell(pfit(1:5))), 'r-', 'LineWidth', 1.5);
title('Endogenous Input Function'); legend('True F^{en}','Estimated F^{en}');